How to properly obtain and use MHFP Fingerprint? #5556
Unanswered
jshan2020-33
asked this question in
Q&A
Replies: 1 comment
-
|
Those are MinHash raw hash values. They are meant to be used with Jaccard similarity, using built-in method for fast comparisons. See: https://iwatobipen.wordpress.com/2018/10/13/new-fingerprint-minhash-fingerprint-rdkit-chemoinformatics/. I'm also looking for a way to use this in generic ML applications. Maybe just use modulo operation? |
Beta Was this translation helpful? Give feedback.
0 replies
Sign up for free
to join this conversation on GitHub.
Already have an account?
Sign in to comment
-
The code used to get MHFP is as follows:
from rdkit import Chem
from rdkit.Chem.rdMHFPFingerprint import MHFPEncoder
mol = Chem.MolFromSmiles('OCC(=O)CCCN')
a = MHFPEncoder()
MHFP = MHFPEncoder.EncodeMol(a, mol)
MHFP_vector = [int(i) for i in MHFP]
If the acquisition method is correct, can MHFP be directly input into the machine learning model as a feature without conversion?
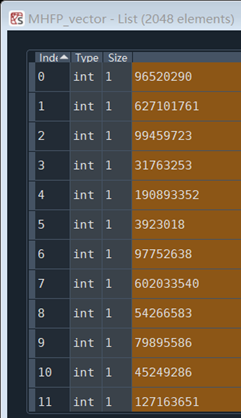
The obtained value of MHFP is not 0/1, but a very large value, as shown in the following figure:
Python: 3.7.0
RDKit: 2020.09.1
Beta Was this translation helpful? Give feedback.
All reactions