You signed in with another tab or window. Reload to refresh your session.You signed out in another tab or window. Reload to refresh your session.You switched accounts on another tab or window. Reload to refresh your session.Dismiss alert
I encountered some cases where CACTVS smiles were wrongly parsed by rdkit while the OpenEye OEToolkits smiles were correctly parsed by rdkit. The cases were for PDB ligands C3O, ZNE, OCC, C33, TX5, O1C (I obtained the smiles from the following PDB web service: https://data.rcsb.org/rest/v1/core/chemcomp/OCC)
Are OpenEye OEToolkits smiles better supported by rdkit than CACTVS smiles?
reacted with thumbs up emoji reacted with thumbs down emoji reacted with laugh emoji reacted with hooray emoji reacted with confused emoji reacted with heart emoji reacted with rocket emoji reacted with eyes emoji
-
Hi,
I encountered some cases where CACTVS smiles were wrongly parsed by rdkit while the OpenEye OEToolkits smiles were correctly parsed by rdkit. The cases were for PDB ligands C3O, ZNE, OCC, C33, TX5, O1C (I obtained the smiles from the following PDB web service: https://data.rcsb.org/rest/v1/core/chemcomp/OCC)
Are OpenEye OEToolkits smiles better supported by rdkit than CACTVS smiles?
Example for ligand OCC:
If I visualise the original smiles in http://www.cheminfo.org/Chemistry/Cheminformatics/Smiles/index.html I can see they are the same:

But after the CACTVS smiles is parsed by rdkit then it becomes a different molecule (the same does not happen with the OpenEye OEToolkits smiles):
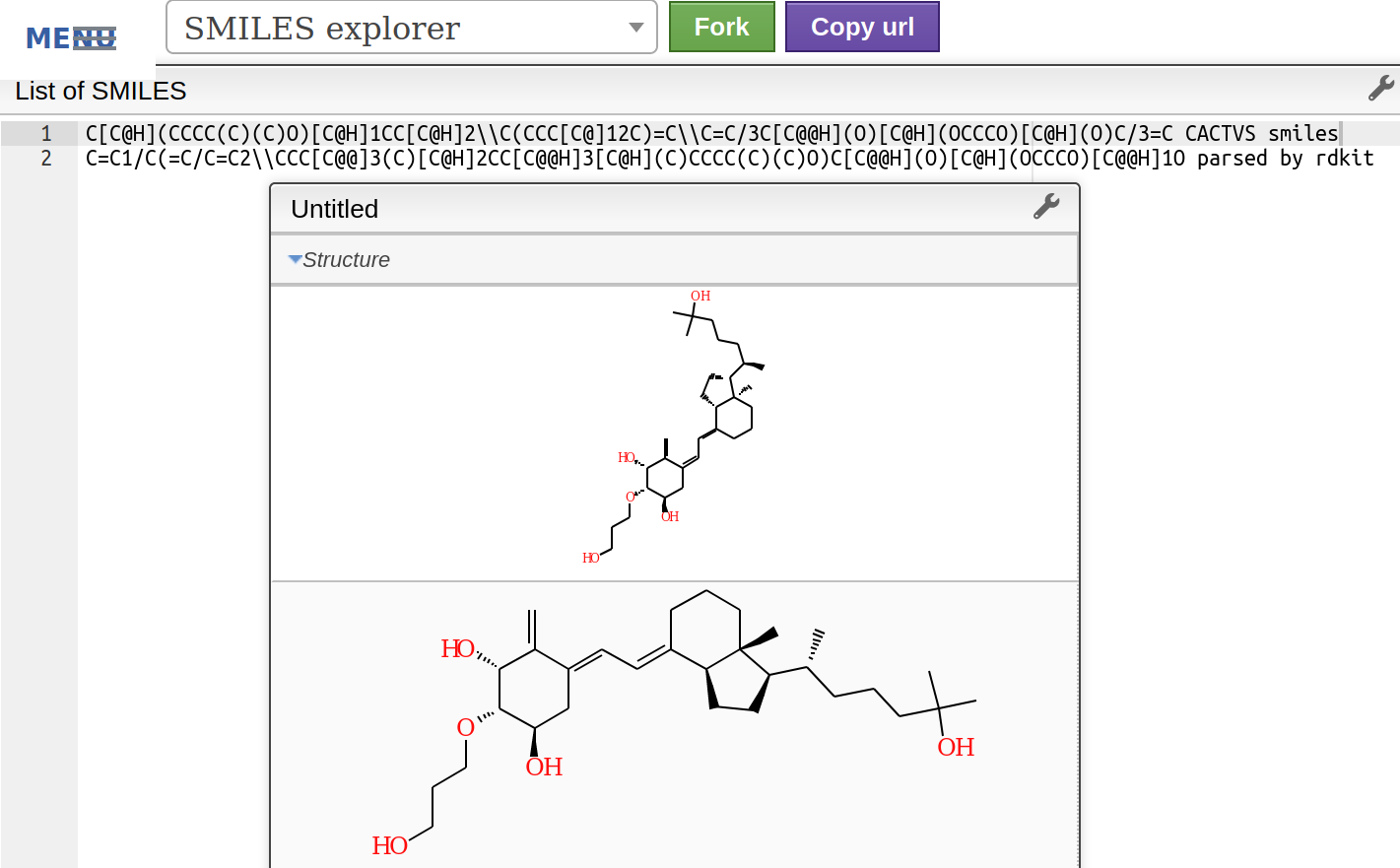
Thank you in advance!
Beta Was this translation helpful? Give feedback.
All reactions