-
Notifications
You must be signed in to change notification settings - Fork 39
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
index out of bounds error with plotting.py #18
Comments
|
This is a problem I haven't run into before! The problem seems to be related to trying to find correlation values for the members of the family in the correlation table, and it isn't finding them (returning an empty array, and then complaining that it has zero length when it expects it to have at least one value). In theory this shouldn't happen. It seems like this is potentially a symptom of a problem that happened earlier in the code that didn't raise an error. A few questions first, and then we can work on a workaround...
I have some ideas to get you back to processing with the hope that this is the only family that has this issue. |
|
Thanks for your comments, and apologies for the late reply, was on leave a couple of days.
Many thanks. |
|
Ah, gotcha! It's possible that something went wrong in the removal step. I haven't run into issues yet, but for now that seems like a possible culprit. For now, my recommendation is this:
Let me know how that goes. |
|
So none of the family plots (starting "fam*.png") rendered? I'm not surprised that the .html didn't render, since it gets rendered last. Let's try |
|
Hmmm, Cluster 0 is the problem, confirmed when |
|
For now, let's have you replace lines 787-792 of redpy/plotting.py with this: Basically, it's going to try to skip trying to find the values of the correlation matrix if it fails. There's probably a more elegant way to do this, but it's difficult to write code without having the problem with me! Pick up where you left off with your processing, and see if the problem persists with the next timestep. I'm going to leave this issue open and not commit these changes in case someone else runs into the issue. I'm not sure what the conditions to reproduce the issue are, and so can't fix the root issue. |
|
Thank you, that work as you intended. I've implemented this as a new local branch for the time being. I agree it's difficult for you when all you have are my 'it doesn't work' comments. And it was really odd that it only occurred after some of my processing and with just this data set. I'd get some more of the data I've been working on and then run REDpy for a further 2-3 weeks of data and let you know how it goes. Probably that won't be until next week. Until then, many thanks. |
|
Great! Also, I found a small issue in the code I sent you. I've edited it so that the "found issue" print message is more informative (variable Good luck! |
|
Thank you. I thought the output of that print was a bit odd, in that it didn't specify family 0, but assumed there was some internal referencing going on. Thanks again! |
|
This is part of the workaround. I put the try/catch around the definition of the variable this is attempting to plot. Since that variable was not filled, it plots it as effectively empty. |
|
Understood. So I'm all good. Thank you for all your help. I have something else that is unrelated, so I'll put it in a separate ticket. |

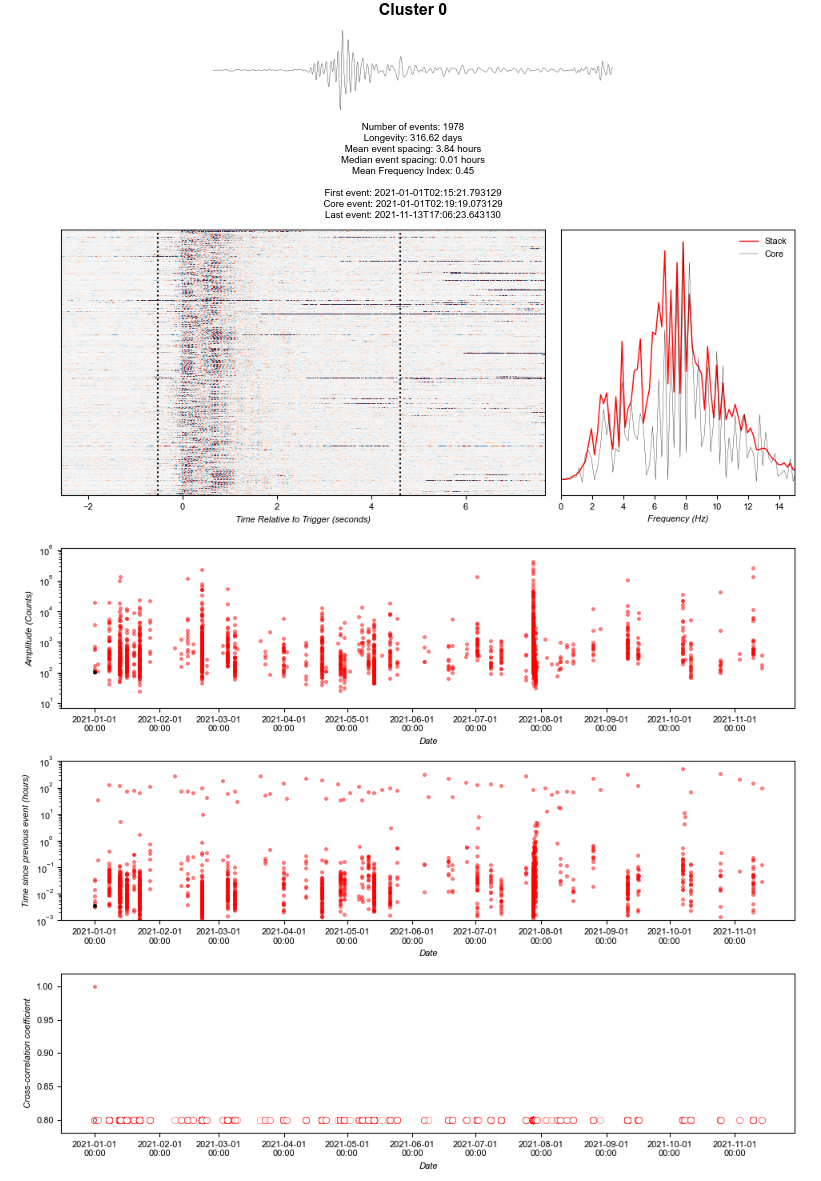
I am running REDpy on a single station using locally stored files. Running Ubuntu 18.04LTS.
I have a total 10 months of data to process. Have run backfill in blocks of a month or two. Up to 4 months all was fine. Then with the 5th, the clustering was okay, but the plotting of clusters failed with an index out of bounds message. This example is from running forcePlot, but the error message is the same.
The relevant code segment is.
I can't see how the plotting is fine until some point and then fails, after which nothing I do seems to get the cluster families to plot again. Could this be something to do with the large number events detected and that clusters have many elements??
Many thanks.
The text was updated successfully, but these errors were encountered: